Protein Structure
We are much focused on the determination of structures via NMR spectroscopy. In the past, bringing proton-detected solid-state NMR forward has been the major focus, where structure calculation based on the newly obtained beneficial properties of protons could be exploited for the first time. We have been interested in innovative proton-proton correlation experiments (e. g., time-shared and higher-dimensionality strategies), paramagnetic tagging, and enzyme complex structure calculation.
Structure calculation characterization of dynamics of biologically interesting via novel tools both in solution and solid-state NMR has become a recent interest of the group.
Past and recent research topics in this direction are, for example:
New spectroscopic approaches for improved structure calculation
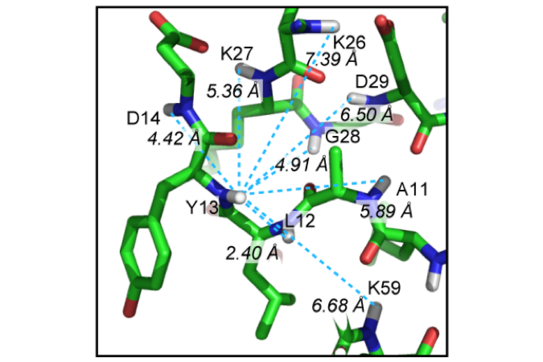
R. Linser, B. Bardiaux, V. Higman, U. Fink, B. Reif,
“Structure Calculation from Unambiguous Long-Range Amide and Methyl 1H−1H Distance Restraints for a Microcrystalline Protein with MAS Solid-State NMR Spectroscopy”,
J. Am. Chem. Soc. 133 (15), 5905–5912 (2011).
R. Linser, B. Bardiaux, S. G. Hyberts, A. Kwan, V. Morris, M. Sunde, G. Wagner,
“Solid-State NMR Structure Determination from Diagonal-Compensated, Sparsely Nonuniform-Sampled 4D Proton–Proton Restraints”,
J. Am. Chem. Soc. 136 (31), 11002–11010 (2014).
Paramagnetic tags for improved structure calculation
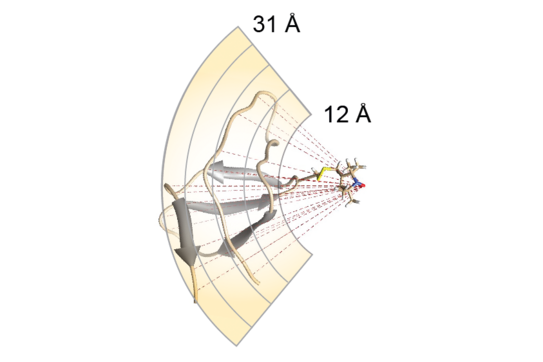
P. Rovó, K. Grohe, K. Giller, S. Becker, R. Linser,
“Proton Transverse Relaxation as a Sensitive Probe for Structure Determination in Solid Proteins”,
ChemPhysChem,, 16 (18), 3791–3796 (2015).
Cover: ChemPhysChem, 16 (18), 3739 (2015).
”Cover Profile” article: ChemPhysChem, 16 (18), 3743 (2015).
Data processing frameworks for improved structure calculation
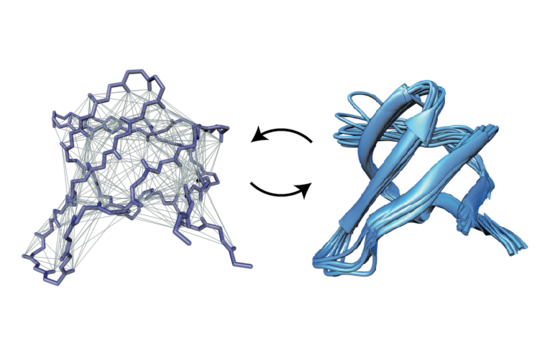
K. Grohe, H. Singh, S. K. Vasa, B. Söldner, E. Nimerovsky, B. Vögeli, C. M. Rienstra, R. Linser,
“Exact distance measurements for structure and dynamics in solid proteins by fast magic angle spinning NMR”,
accepted in Chem. Commun. (2019).
Structure determination of in the absence of deuteration
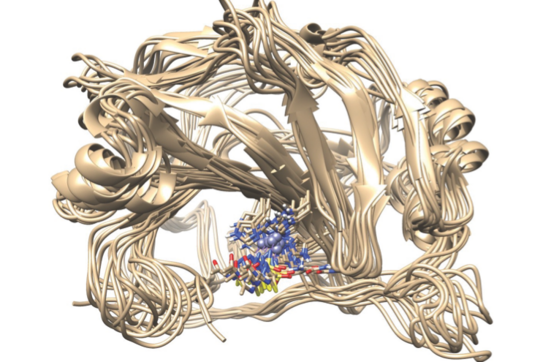
S. K. Vasa, H. Singh, K.Grohe, R. Linser,
"Assessment of a large enzyme-drug complex by proton-detected solid-state NMR without deuteration",
Angew. Chem. Int. Ed., DOI: 10.1002/anie.201811714 (2019).
Cover article
Back





